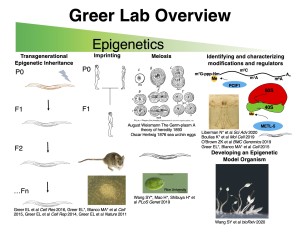
The Greer lab is broadly interested in the molecular mechanisms of epigenetics. We feel that the most exciting scientific discoveries occur at the interface of multiple fields and therefore strive to tackle long-standing fundamental questions from new angles. We address questions using a combination of genetic/genomic, molecular, cellular, and biochemical approaches in a variety of model organisms including D. discodieum, C. elegans, M. musculus, and human cell lines. This philosophy has brought our lab into many exciting fields including, longevity, DNA damage and repair, virology, ribosome heterogeneity, epitranscritomics, and the evolution of multicellular organisms.
In its strictest definition, epigenetics encompasses the study of heritable non-genetic information, In this strict sense of the term, we study transgenerational epigenetic inheritance – non-genetic information being transmitted from ancestors to their descendants for multiple generations to regulate complex biological phenomena. In more recent years, the field of epigenetics has expanded to include the study of how epigenetic information, such as modifications to chromatin and DNA, regulate a variety of cellular processes. A portion of the lab is focused on identifying new epigenetic modifications and the enzymes that add, remove, and recognize these modifications. We examine how these enzymes and modifications regulate biological processes such as the viral response, meiosis, stress resistance, and aging . Finally a portion of the lab is repurposing the slime mold Dictyostelium discodieum as a new model organism to examine the role of epigenetics in the evolution of multicellularity. In the sections below, we will briefly describe some of our forays into each of these layers of epigenetic regulation.
Transgenerational Epigenetic Inheritance
We have previously identified chromatin-modifying enzymes that regulate complex transgenerational phenotypes in C. elegans, including longevity and fertility. Mutation of a histone H3 lysine 4 (H3K4) trimethylation complex regulates worm lifespan in both the generation in which the mutation occurs and in subsequent generations lacking the mutation. More recent work focuses on understanding how deletion of the C. elegans H3K4me2 demethylase, spr-5, leads to inherited accumulation of the euchromatic H3K4me2 mark and a progressive decline in fertility in successive generations. To investigate the underlying molecular mechanism behind progressive sterility of spr-5 mutant worms, we carried out an RNAi screen and identified both suppressors and enhancers of sterility. This screen led to a working model for how epigenetic changes in histone H3 methylation might be inherited to affect complex organismal traits. Current projects are focused on identifying new paradigms of epigenetic inheritance and characterization of which epigenetic cues are transmitted from ancestors to their descendants to regulate complex biological phenotypes. We have recently performed metabolic methyl labeling experiments to track heritable methylation from parents to their children and found that in response to parental starvation there is an increase in 18S rRNA methylation that is passed to children to alter translation in preparation for less food availability. Parental starvation leads to a hormesis phenotype, where one stress will induce the stress response pathways so that organisms respond better to other stresses. In C. elegans, parental starvation leads to increased heat stress resistance, a subtle increase in lifespan, and a reduction in fertility that is passed on to the children and grandchildren before returning to basal levels in the F3 generation. We find that deletion of the 18S rRNA methyltransferases eliminates the transmission of this transgenerational hormesis phenotypes to the naive well fed descendants.
Identification and characterization of new epigenetic modifications
We seek to identify new epigenetic and epitranscriptomic modifications, the enzymes which regulate these modifications, and their role in a variety of biological processes. Current projects are focused on how epigenetic modifications can regulate cancer metastasis, longevity, and stress resistance in a variety of model organisms. For example, we have identified PCIF1 as an mRNA methyltransferase which adds a methyl group to adenosines on the cap of mRNAs in mice and humans to produce a dually methylated adenosine (m6Am). Our current projects examine the biological function of this capping mechanism in processes such as viral replication and neural development. We also identified the enzyme METL-5 as a m6A methyltransferase which methylates a single residue on the 18S ribosomal rRNA to regulate lipid oxidation and stress resistance in C. elegans. Our identification of METL-5 as an 18S rRNA methyltransferase has taken us into the field of ribosome heterogeneity and lipid signaling. Click here for a recent talk Eric gave on PCIF1 and METL-5
Dictyostelium discodieum
The evolutionary transition from unicellular to multicellular organisms remains a fundamental mystery of biology. The explosion of epigenetic complexity at this transitional time point led us to hypothesize that epigenetic changes helped drive the evolution of multicellularity. We are examining this hypothesis in Dictyostelium discodieum, one of the rare organisms which is both unicellular and multicellular. We recently performed a comprehensive profiling of chromatin state and single cell gene expression in Dictyostelium at both its unicellular and multicellular stages. Unlike the unidirectional development from a unicellular zygote into a multicellular organism, Dictyostelium readily transits back and forth between both differentiated multicellular and unicellular states. By comparing the epigenome of Dictyostelium at both unicellular and multicellular stages to syntenic regions in the unicellular eukaryote S. pombe and the multicellular eukaryote C. elegans, we identify a conserved multicellular chromatin accessibility signature. This work helped to identify specific, conserved epigenetic modifications that facilitate the evolution of multicellularity and demonstrated a critical role for epigenetics in multicellularity. Dictyostelium affords us the capacity to manipulate the genome and epigenome of this organism to identify genes as well as epigenetic modifications that are necessary and sufficient for multicellularity in Dictyostelium. We are currently using Dictyostelium to examine the role of epigenetics in the evolution of cellular altruism and epigenetic memory. Click here for a recent talk Eric gave on METL-5 and Dictyostelium.
Videos from David Knecht and Rex Chisholm Labs